
Bayesian Evolutionary Layered Learning Architectures (BELLA) is a BEAST 2 package that brings unsupervised Bayesian neural networks to phylodynamic inference, letting you map rich predictor sets (traits, time series, environmental covariates) to key parameters such as speciation, extinction, transmission, and migration rates — all learned jointly with the phylogeny via MCMC.
🧭 Overview
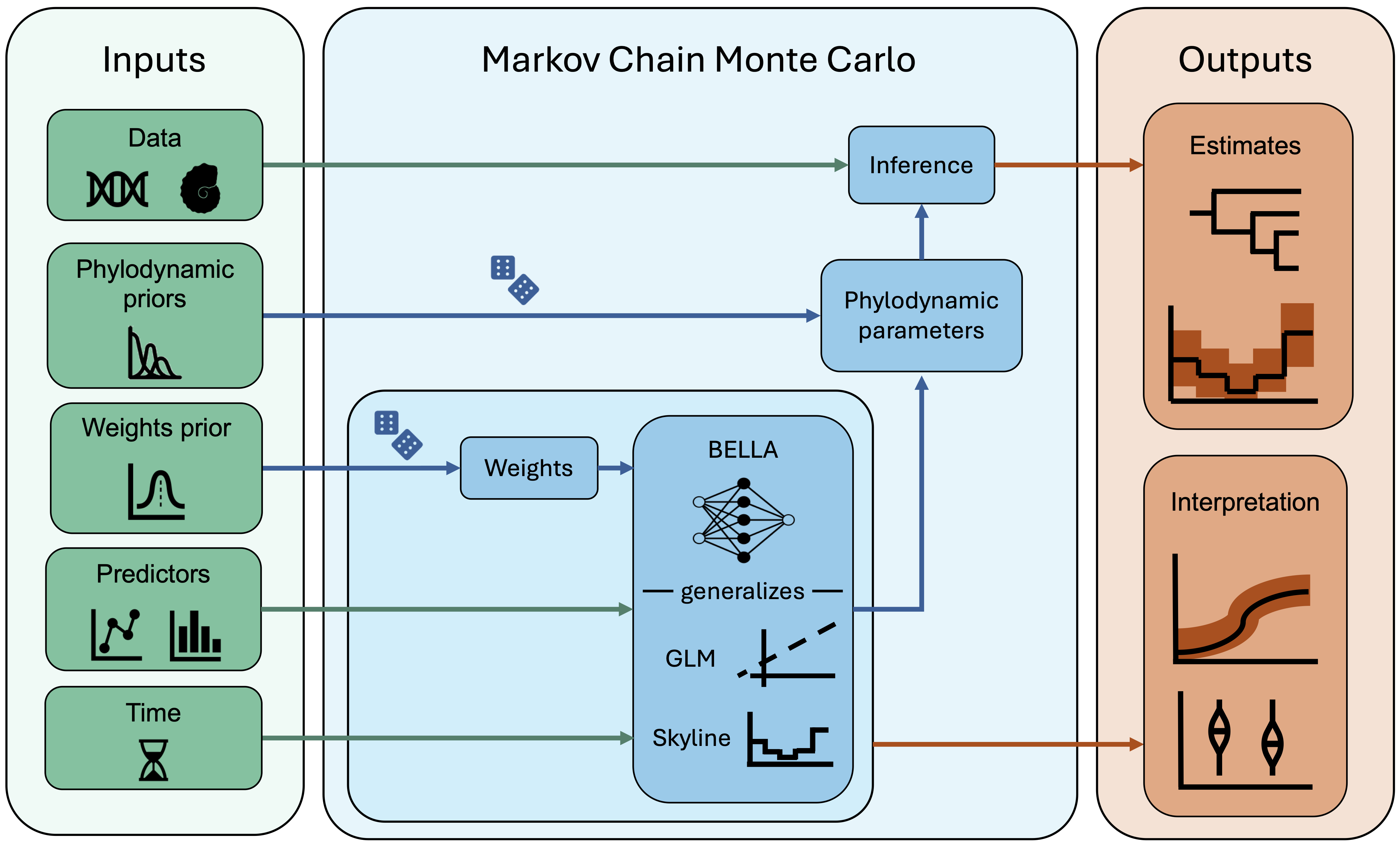
BELLA integrates phylogenetic data with predictor covariates—such as traits, environmental time series, or mobility patterns—and prior knowledge on phylodynamic parameters. Importantly, this is formulated as an unsupervised learning problem: no evolutionary or epidemiological rates are observed or used as training targets. Instead, BELLA employs a Bayesian neural network to flexibly learn how predictors relate to key parameters—such as speciation, extinction, transmission, or migration rates—directly through the phylogenetic likelihood. This allows the model to capture complex, nonlinear dependencies that go beyond traditional generalized linear models (GLMs) and skyline models. All parameters, including the neural network weights, are estimated jointly within a Markov chain Monte Carlo framework. The resulting posterior distributions enable both inference of population dynamics and interpretation of predictor effects using explainable AI tools.
✨ Features
- 🧠📈 Nonlinear, highly flexible mappings that model smooth, high-dimensional relationships between predictors and phylodynamic parameters, generalizing and extending GLMs and skyline models without restrictive linearity or piecewise-constant assumptions.
- 🧬🤖 Unsupervised Bayesian neural networks whose weights are inferred jointly with the phylogeny and evolutionary parameters by conditioning directly on the sequence likelihood—eliminating the need for external response variables or labeled training data.
- ⚖️🔒 Built-in regularization via weight priors that penalize overly complex functions, reducing overfitting while propagating uncertainty into phylodynamic estimates.
- 🔍🧩 Explainable AI tooling using partial dependence plots and SHAP-style attribution scores to quantify and visualize predictor effects on inferred phylodynamic parameters.
- ⚙️🐒🦠 Deep BEAST 2 integration enabling end-to-end Bayesian inference for epidemiological and macroevolutionary workflows within existing BEAST 2 pipelines.
⚙️ Installation
BELLA is available as a package for BEAST 2.7+ . You can install it via BEAUti as follows:
- Open BEAUti.
- Go to
File→Manage Packages. - Click
Package repositories. Add URLand enter:https://raw.githubusercontent.com/gabriele-marino/BELLA/main/package.xml.- Click
OKto add the repository. - Close the Package Repositories window. Return to the Package Manager window, scroll down to find
BELLA, select it, and click Install/Upgrade. - You're now ready to use BELLA!
🚀 Getting started
We provide several well-documented example configuration files in the BELLA examples directory, covering a range of use cases. See the examples README to get started!
If you are new to BEAST 2, consider exploring the tutorials to familiarize yourself with BEAST XML configuration files and MCMC analyses.
If you’ve already run your analyses and want to postprocess the results, check out the BELLA-companion Python package, which includes helpful plotting tools like the ones shown above.
❗️ If you don't find what you're looking for, please reach out—I’m always happy to help!
📑 Citing BELLA
If you use BELLA in your research, please cite the following:
COMING SOON
📫 Contact
For questions, bug reports, or feature requests, please, consider opening an issue on GitHub, or contact me directly.
For help with configuration files, don’t hesitate to reach out — I’m happy to assist!